
Pathway Design through Machine Learning
Leverage machine learning and probabilistic modeling to guide experiments
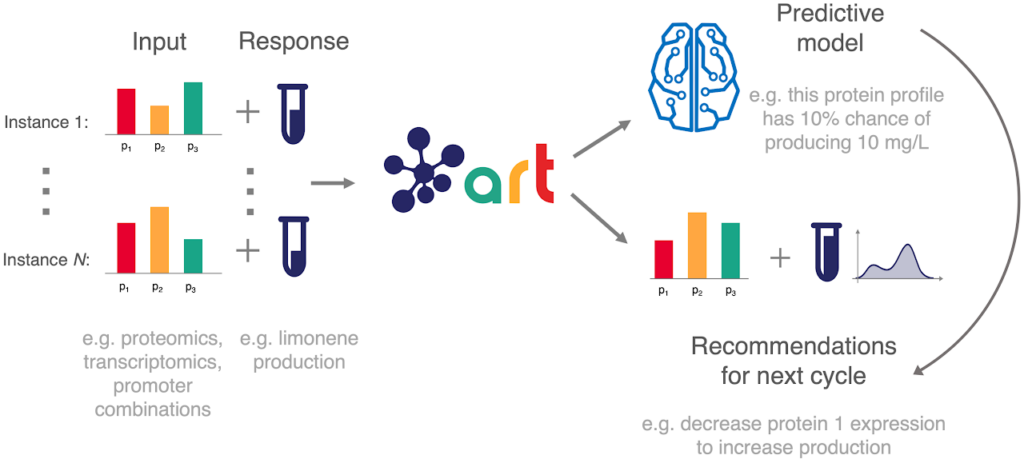
Our machine learning algorithm — ART: the Automated Recommendation Tool — leverages machine learning and probabilistic modeling techniques to guide synthetic biology in a systematic fashion, without the need for a full understanding of the biological system. It provides a predictive model and recommendations for the next cycle.
ART has been effectively demonstrated on both simulated and actual experimental data from metabolic engineering projects producing a variety of bioproducts. ART is both intuitive and easy to use, lowering the barrier to machine learning access.
Publications
- A machine learning Automated Recommendation Tool for synthetic biology
- Combining mechanistic and machine learning models for predictive engineering and optimization of tryptophan metabolism