
By Aliyah Kovner
By modifying the genomes of plants and microorganisms, synthetic biologists can design biological systems that meet a specification, such as producing valuable chemical compounds, making bacteria sensitive to light, or programming bacterial cells to invade cancer cells. This field of science, though only a few decades old, has enabled large-scale production of medical drugs and established the ability to manufacture petroleum-free chemicals, fuels, and materials. It seems that biomanufactured products are here to stay, and that we will rely on them more and more as we shift away from traditional, carbon-intensive manufacturing processes.
But there is one big hurdle – synthetic biology is labor intensive and slow. From understanding the genes required to make a product, to getting them to function properly in a host organism, and finally to making that organism thrive in a large-scale industrial environment so it can churn out enough product to meet market demand, the development of a biomanufacturing process can take many years and many millions of dollars of investment.
Héctor García Martín, a staff scientist in the Biosciences Area of Lawrence Berkeley National Laboratory (Berkeley Lab) and the Agile BioFoundry, is working to accelerate and refine this R&D landscape by applying artificial intelligence and the mathematical tools he mastered during his training as a physicist.
We spoke with him to learn how AI, bespoke algorithms, mathematical modeling, and robotic automation can come together as a sum greater than its parts, and provide a new approach for synthetic biology.
Q. Why do synthetic biology research and process scale-up still take a long time?
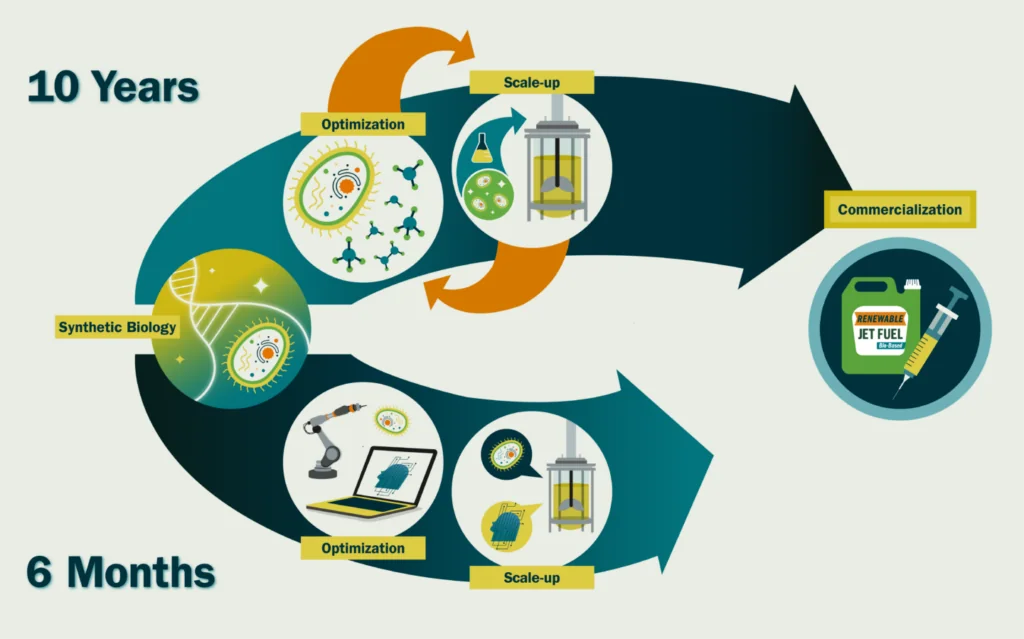
García Martín: I think the hurdles we find in synthetic biology to create renewable products all stem from a very fundamental scientific shortcoming: our inability to predict biological systems. Many synthetic biologists might disagree with me and point towards the difficulty in scaling processes from milliliters to thousands of liters, or the struggles to extract high enough yields to guarantee commercial viability, or even the arduous literature searches for molecules with the right properties to synthesize. And, that is all true. But I believe they are all a consequence of our inability to predict biological systems. Say we had someone with a time machine (or God, or your favorite omniscient being) come and give us a perfectly designed DNA sequence to put in a microbe so it would create the optimal amount of our desired target molecule (e.g, a biofuel) at large scales (thousands of liters). It would take a couple of weeks to synthesize and transform it into a cell, and three to six months to grow it at a commercial scale. The difference between those 6.5 months and the ~10 years that it takes us now, is time spent fine tuning genetic sequences and culture conditions – for example, lowering expression of a certain gene to avoid toxic build-up or increasing oxygen levels for faster growth – because we don’t know how these will affect cell behavior. If we could accurately predict that, we would be able to engineer them much more efficiently. And that is how it is done in other disciplines. We do not design planes by building new plane shapes and flying them to see how well they work. Our knowledge of fluid dynamics and structural engineering is so good we can simulate and predict the effect that something like a fuselage change will have on flight.
Q. How does artificial intelligence accelerate these processes? Can you give some examples of recent work?
García Martín: We are using machine learning and artificial intelligence to provide the predictive power that synthetic biology needs. Our approach bypasses the need to fully understand the molecular mechanisms involved, which is how it saves significant time. However, this does raise some suspicion in traditional molecular biologists.
Normally these tools have to be trained on huge datasets, but we just don’t have as much data in synthetic biology as you might have in something like astronomy, so we developed unique methods to overcome that limitation. For example, we have used machine learning to predict which promoters (DNA sequences that mediate gene expression) to choose to get maximum productivity. We have also used machine learning to predict the right growth media for optimal production, to predict metabolic dynamics of cells, to increase the yields of sustainable aviation fuel precursors, and to predict how to engineer functioning polyketide synthases (enzymes that can produce an enormous variety of valuable molecules but are infamously difficult to predictably engineer).
In many of these cases we needed to automate the scientific experiments to obtain the large amounts of high-quality data that we need for AI methods to be truly effective. For example, we have used robotic liquid handlers to create new growth media for microbes and test their effectiveness, and we have developed microfluidic chips to try to automate genetic editing. I am actively working with others at the Lab (and external collaborators) to create self-driving labs for synthetic biology.
Q. Are many other groups in the U.S. doing similar work? Do you think this field will get bigger in time?
The number of research groups with expertise in the intersection of AI, synthetic biology, and automation is very small, particularly outside of industry. I would highlight Philip Romero at the University of Wisconsin and Huimin Zhao at the University of Illinois Urbana-Champaign. However, given the potential of this combination of technologies to have a huge societal impact (e.g., in combating climate change, or producing novel therapeutic drugs), I think this field will grow very fast in the near future. I have been part of several working groups, commissions, and workshops, including a meeting of experts for the National Security Commission on Emerging Biotechnology, that discussed the opportunities in this area and are drafting reports with active recommendations.
Q. What kind of advances do you anticipate in the future from continuing this work?
García Martín: I think an intense application of AI and robotics/automation to synthetic biology can accelerate synthetic biology timelines ~20-fold. We could create a new commercially viable molecule in ~6 months instead of ~10 years. This is direly needed if we want to enable a circular bioeconomy – the sustainable use of renewable biomass (carbon sources) to generate energy and intermediate and final products.
There are an estimated 3,574 high‐production‐volume (HPV) chemicals (chemicals that the U.S. produces or imports in quantities of at least 1 million pounds per year) that come from petrochemicals nowadays. A biotechnology company called Genencor needed 575 person-years of work to produce a renewable route for producing one of these widely used chemicals, 1,3-propanediol, and this is a typical figure. If we assume that’s how long it would take to design a biomanufacturing process to replace the petroleum-refining process for each of these thousands of chemicals, we would need ~2,000,000 person years. If we put all the estimated ~5,000 U.S. synthetic biologists (let’s say 10% of all biological scientists in the U.S., and that is an overestimate) to work on this, it would take ~371 years to create that circular bioeconomy. With the temperature anomaly growing every year, we don’t really have 371 years. These numbers are obviously quick back-of-the envelope calculations, but they give an idea of the order of magnitude if we continue the current path. We need a disruptive approach.
Furthermore, this approach would enable the pursuit of more ambitious goals that are unfeasible with current approaches, such as: engineering microbial communities for environmental purposes and human health, biomaterials, bioengineered tissues, etc.
Q. How is Berkeley Lab a unique environment to do this research?

García Martín: Berkeley Lab has had a strong investment in synthetic biology for the last two decades, and displays considerable expertise in the field. Moreover, Berkeley Lab is the home of “big science”: large-team, multidisciplinary science, and I think that is the right path for synthetic biology at this moment. Much has been achieved in the last seventy years since the discovery of DNA through single-researcher traditional molecular biology approaches, but I think the challenges ahead require a multidisciplinary approach involving synthetic biologists, mathematicians, electrical engineers, computer scientists, molecular biologists, chemical engineers, etc. I think Berkeley Lab should be the natural place for that kind of work.
Q. Tell us a little about your background, what inspired you to study mathematical modeling of biological systems?
García Martín: Since very early on, I was very interested in science, specifically biology and physics. I vividly remember my dad telling me about the extinction of dinosaurs. I also remember being told how, in the Permian period, there were giant dragonflies (~75 cm) because oxygen levels were much higher than now (~30% vs 20%) and insects get their oxygen through diffusion, not lungs. Hence, larger oxygen levels enabled much larger insects. I was also fascinated by the ability that mathematics and physics afford us in understanding and engineering things around us. Physics was my first choice, because the way biology was taught in those times involved a lot more memorization than quantitative predictions. But I always had an interest in learning what scientific principles led to life on Earth as we see it now.
I obtained my Ph.D. in theoretical physics, in which I simulated Bose-Einstein condensates (a state of matter that arises when particles called bosons, a group that includes photons, are at close to absolute zero temperature) and using path integral Monte Carlo techniques, but it also provided an explanation for a 100+ year old puzzle in ecology: why does the number of species in an area scale with an apparently universal power law dependence on the area (S=cAz, z=0.25)? From then on I could have continued working on physics, but I thought I could make more of an impact by applying predictive capabilities to biology. For this reason, I took a big gamble for a physics Ph.D. and accepted a postdoc at the DOE Joint Genome Institute in metagenomics – sequencing microbial communities to unravel their underlying cellular activities – with the hope of developing predictive models for microbiomes. I found out, however, that most microbial ecologists had limited interest in predictive models, so I started working in synthetic biology, which needs predictions capabilities because it aims to engineer cells to a specification. My current position allows me to use my mathematical knowledge to try and predictably engineer cells to produce biofuels and fight climate change. We have made a lot of progress, and have provided some of the first examples of AI-guided synthetic biology, but there is still a lot of work to do to make biology predictable.
This story originally appeared on the Berkeley Lab News Center.